Catching the Risk: The Maths Behind Sample Quotas in SEGs
In occupational hygiene, we often talk about Similar Exposure Groups (SEGs) and the need to collect enough samples to be confident in our data. But how do we actually determine if our sample size is sufficient to capture those at the highest risk?
Standard practice is to mornally just quote Technical Appendix A in NIOSH 173/77 and go from there – but this time, that just wasn’t going to be enough.
During a recent mentoring session, we broke down the statistics of sampling quotas. Here is how the maths of permutations and combinations helps us ensure no worker is left behind.
The Foundation: Factorials and Choices
Before we can calculate the probability of a successful sampling campaign, we need to understand how many ways we can “choose” workers from a group.
Factorials
The factorial ($n!$) is the product of all positive integers less than or equal to $n$. This value represents the total number of ways you can uniquely arrange a set of items. For example, if you have 6 items and want to know every possible sequence they could be ordered in:
$$6! = 6 \times 5 \times 4 \times 3 \times 2 \times 1 = \mathbf{720 \text{ ways to arrange 6 items}}$$
Permutations vs. Combinations
Understanding the difference between these two is critical for sampling.
Permutations
How many ways can I choose $r$ things from $n$ things where order is important? To use our example of 6 items, let’s say I want to select 2. I can do that by:
$$6 \times 5 = 30$$
To get that, using only factorials, we can use some numerator/denominator cancellation. It looks like this, noting how $4 \times 3 \times 2 \times 1$ appears in both the top and bottom and so cancel out:
$$^nP_r = \frac{6!}{(6-2)!} = \frac{6!}{4!} = \frac{6 \times 5 \times 4 \times 3 \times 2 \times 1}{4 \times 3 \times 2 \times 1} = 6 \times 5 = 30$$
MAGIC!
The general formula is:
$$^nP_r = \frac{n!}{(n-r)!}$$
Combinations
How many ways can I choose $r$ things from $n$ things where order is NOT important? In our example of 30 permutations of 2 items, we divide by the 2 ways those items can be arranged ($2 \times 1 = 2$). So let’s divide our permutation formula by the factorial of $r$:
$$\frac{6!}{2! \times (6-2!)} = \frac{6 \times 5 \times 4 \times 3 \times 2 \times 1}{(2 \times 1)(4 \times 3 \times 2 \times 1)} = \frac{6 \times 5}{2} = 15$$
$$^n\mathcal{C}_r = \frac{n!}{r! \times (n-r)!}$$
Practical Example: Choosing 3 Workers from 10
To see why this distinction matters, imagine you have a group of 10 workers ($n=10$) and you want to select 3 ($r=3$) for monitoring.
The Permutation Approach (Order Matters)
If we cared about the order (e.g., who gets the pump first, second, and third), the maths looks like this:
$$^{10}P_3 = \frac{10!}{(10-3)!} = \frac{10!}{7!} = 10 \times 9 \times 8 = \mathbf{720 \text{ ways}}$$
The Combination Approach (Order Does Not Matter)
In a typical hygiene survey, we don’t care about the sequence—only the final set of people monitored. We divide the permutations by the ways those 3 people can be rearranged ($3!$):
$$^{10}\mathcal{C}_3 = \frac{720}{3 \times 2 \times 1} = \frac{720}{6} = \mathbf{120 \text{ ways}}$$
Why this matters: There are far fewer unique “groups” than “orders.” We use combinations because we are looking for the probability of a specific subset of the population being captured but what order we select them is not important.
Defining the Sampling Problem
When sampling a population ($N$), we are often looking for a specific “Target” group ($N_0$), such as a Highly Exposed Risk Group (e.g., the top 10% of exposed workers).
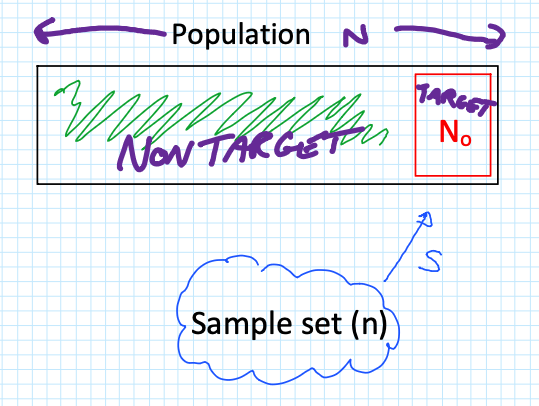
The critical question is: What is the probability that I get ZERO samples from my high-risk group?
To find this, we look at the three components of the probability calculation:
Combinations of $n$ samples from the non-target group $(N – N_0)$.
Combinations of 0 samples from the target group $(N_0)$.
Total possible combinations of $n$ samples from the entire population ($N$).
The maths behind this derivation follows a logical flow:
$$P(r=0) = \frac{^{(N-N_0)}\mathcal{C}_n \times ^{N_0}\mathcal{C}_0}{^N\mathcal{C}_n}$$
Since any combination of 0 items ($^{N_0}\mathcal{C}_0$) is equal to 1 (there is only 1 way to select nothing; it makes maths-sense, go with it), we expand the factorials as follows:
$$P(r=0) = \frac{(N-N_0)!}{n!(N-N_0-n)!} \times \frac{N_0!}{0!(N_0-0)!} \times \left[ \frac{N!}{n!(N-n)!} \right]^{-1}$$
By simplifying the expressions (flipping the $^N\mathcal{C}_n$ with a $-1$ power) and cancelling out terms (specifically $n!$), we arrive at the final simplified formula:
$$P(r=0) = \frac{(N-N_0)!}{(N-N_0-n)!} \times \frac{(N-n)!}{N!}$$
Case Study: Sampling a Population of 18
Let’s apply this to a group of 18 workers ($N=18$). We want to ensure we haven’t missed the top 10% of exposed workers ($18 \times 0.1 = 1.8$, which we call 2 workers, so $N_0 = 2$).
If we collect different sample sizes ($n$), how does your risk of “missing” ($r=0$) that high-risk group change?
Scenario A: Collect 9 Samples ($n=9$)
Using our formula $\frac{(N-N_0)!}{(N-N_0-n)!} \times \frac{(N-n)!}{N!}$:
$$P(r=0) = \frac{(18-2)!}{(18-2-9)!} \times \frac{(18-9)!}{18!} = \frac{16!}{7!} \times \frac{9!}{18!} = \mathbf{0.235}$$
With 9 samples, there is a 23.5% chance you will not capture a single person from that top 10% high-risk group.
Scenario B: Collect 12 Samples ($n=12$)
$$P(r=0) = \frac{(18-2)!}{(18-2-12)!} \times \frac{(18-12)!}{18!} = \frac{16!}{4!} \times \frac{6!}{18!} = \mathbf{0.098}$$
By increasing to 12 samples, your risk of missing the high-risk group drops to roughly 9.8%.
Scenario C: Collect 15 Samples ($n=15$)
$$P(r=0) = \frac{(18-2)!}{(18-2-15)!} \times \frac{(18-15)!}{18!} = \frac{16!}{1!} \times \frac{3!}{18!} = \mathbf{0.0196}$$
At 15 samples, the probability of missing the target group is less than 2%, providing a very high level of certainty.
The “Flip It” Rule
In hygiene, we usually want to know the confidence level—the probability of getting at least one sample from the target group ($r \ge 1$). To find this, simply “flip” the result:
$$\text{Confidence} = 1 – P(r=0)$$
With 12 samples, you have a 90.2% confidence level
i.e. $(1-0.098) \times 100$.
With 15 samples, you have a 98.04% confidence level
i.e. $(1-0.0196) \times 100$.
By using these calculations, you can move away from “guessing” your sample quotas and start providing a statistical justification for your monitoring programs.
BONUS: And I made a calculator for you!!